Correlation
In R, basic correlation tests are executed with two commands: cor()
and lm() (where lm stands for linear model).
Calculating correlation
To calculate product moment correlation coefficient between Maxle and
Maxwi for bronze spears:
> Bronze = subset(spearhead, subset=Mat=="1")
> cor(Bronze$Maxle, Bronze$Maxwi)
[1] 0.6892216
To calculate Spearman's rank correlation coefficient between Date and
Weight for bronze spears:
> cor(Bronze$Date, Bronze$Weight, method="spearman")
[1] 0.1269293
Plotting correlation
To draw a scatterplot for Maxle and Maxwi:
> plot(Bronze$Maxle, Bronze$Maxwi)
The scatterplot by itself is already interesting, but R gives us another
interesting function with the lm() command (where lm stands for
linear model).
> result <- lm(Bronze$Maxwi ~ Bronze$Maxle)
> result
Call:
lm(formula = Maxwi ~ Maxle)
Coefficients:
(Intercept) Maxle
1.5053 0.1277
- note that the order of arguments to
lm()is inverse: the basic use islm(y ~ x)(withyas dependent variable) - the result of
lm()is a rect. You can see by yourself plotting it over the scatterplot
> abline(result$coefficients, col="blue")
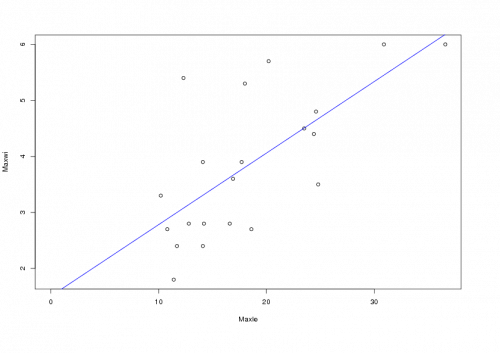
Plotting the lm() result by itself like
> plot(result)
gives you more informative graphs about the linear model, but their content is beyond the scope of this tutorial.